Part:BBa_K4275012
CipA1B2C
CipA1B2C, a cellulosome integrating protein A with one integral carbohydrate-binding module 3 (CBM3), two type I cohesins, and a type II dockerin, is the primary scaffoldin of a cellulosome complex. The CBM3 binds to a range of polysaccharides, including cellulose and PET, increasing the catalytic efficiency of the enzymes. Meanwhile, the type I cohesins interact with the type I dockerins of cellulosomal enzymes, enabling the enzymes to attach to CipA1B2C. CipA1B2C forms a complete cellulosome complex when it binds to outer layer protein B (OlpB), an anchoring protein attached to a cell surface[1]. CipA1B2C improves the efficiency of cellulosomal enzymes by increasing binding affinity to cellulose and PET through CBM and facilitating the spatial proximity among various enzymes, which allows enzymes to work in synergy. The ability to bind to various cellulosomal enzymes makes CipA a versatile tool for enhancing the efficiency of enzymes and conducting different enzyme synergies, which means CipA can promote the degradation rate of diverse materials (i.e. PET and cellulose)[1]. This is a part in a part collection where we enable efficient degradation of cellulose and PET in textile waste.
Figure 1 The 3D structure of the protein predicted by Alphafold2.
Usage and Biology
CipA1B2C is a scaffold protein with one carbohydrate-binding module 3 (CBM3) and two type I cohesins. Natural CipA is found in Clostridium thermocellum, a gram-positive thermophilic and anaerobic bacterium. Cellulosomal enzymes bind tightly to CipA1B2C via the high-affinity dockerin I: cohesin I noncovalent interaction. CipA1B2C is then attached to OlpB, the secondary scaffoldin of the cellulosome complex, through the interaction between the type II dockerin and type II cohesin. OlpB is anchored to the cell surface. The CipA1B2C, OlpB, and the cellulosomal enzymes together comprise a multiplex cellulosome. Utilizing the highly effective cellulosome complex, C.thermocellum is the most efficient microorganism for lignocellulosic biomass degradation.
Characterisation of the Cellulosome Complex
Mini-scaffold construction of cellulosome
We constructed E.coli expression vectors for the mini-scaffold protein subunits. The scaffoldin components of the wild-type cellulosome subunits are large protein scaffolds that would bring a massive protein burden to the bacterial host secreting them. We modified the coding sequences for the wild-type cellulosome protein scaffold, as shown in (Fig.2A and Fig.2B) The mini-scaffolds were successfully expressed by our host, verified by the SDS-PAGE analysis shown in (Fig.2C and Fig.2D)
Functionality testing of our mini-scaffold
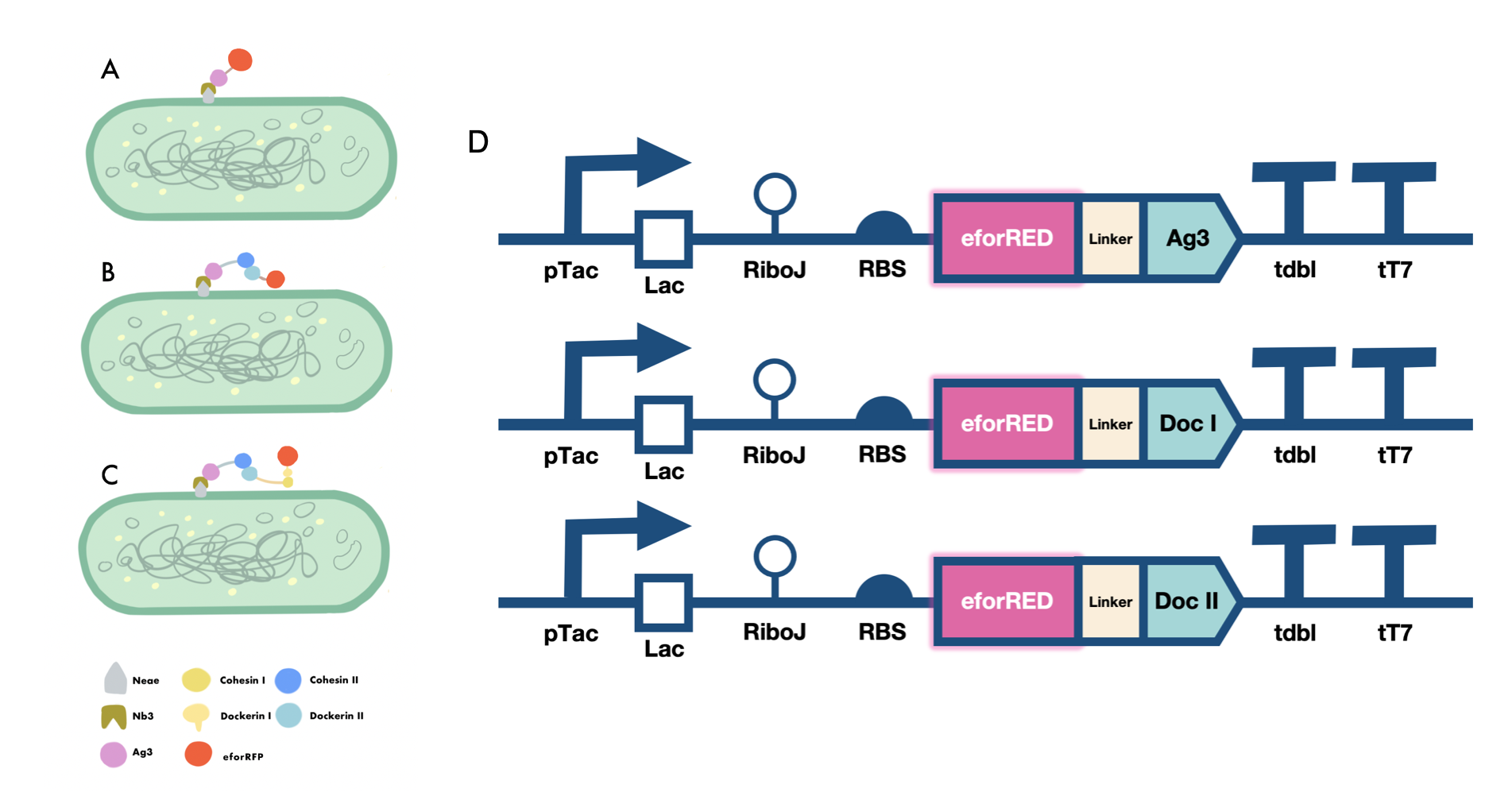
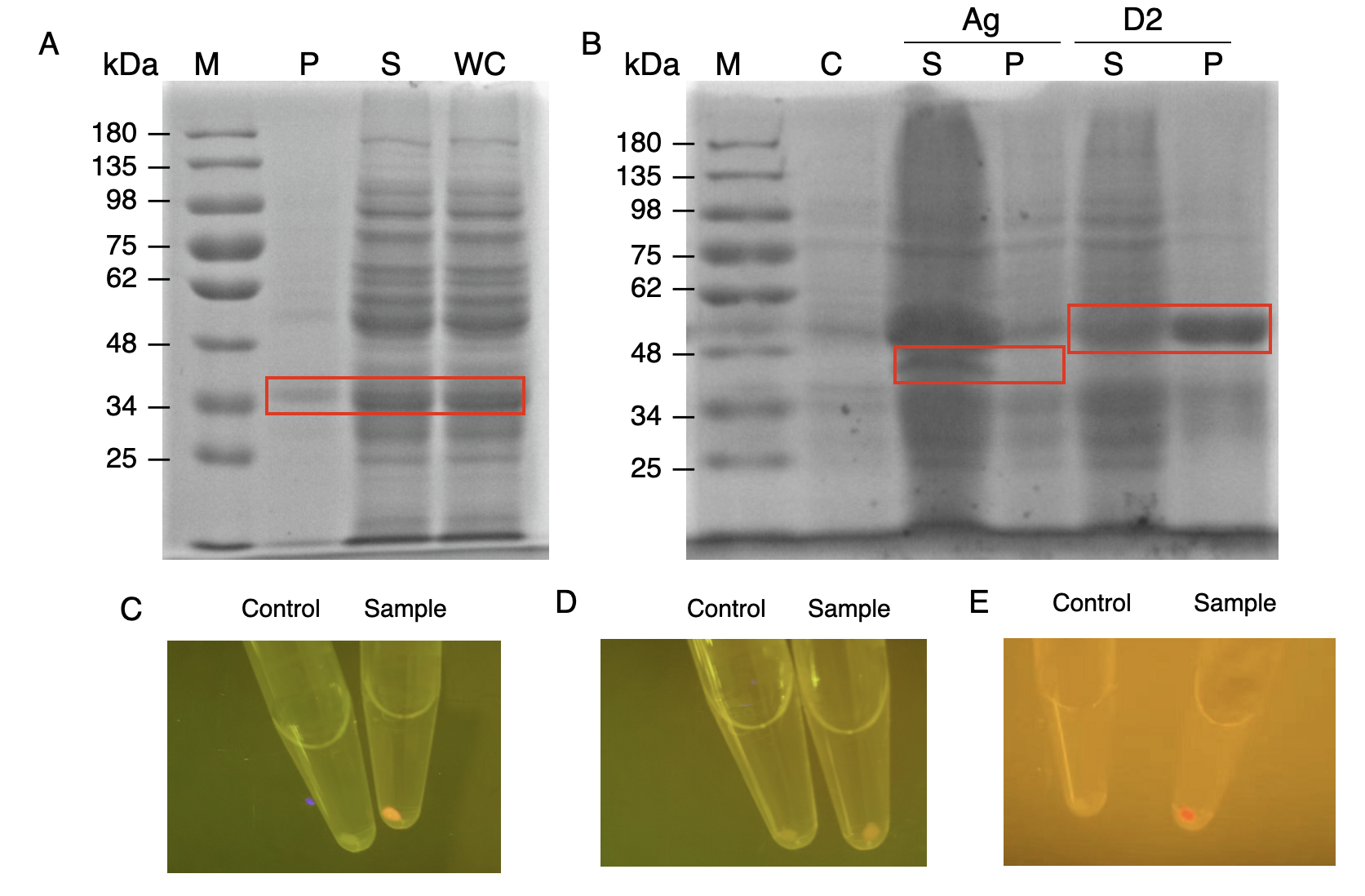
In order to verify the three levels of protein-protein interaction that assembles our cellulosome complex, Ag3-eforRED, DocI-eforRED, and DocII-eforRED vectors were constructed and cultured for IPTG-inducible expression (Fig. 3D). SDS-page analysis was performed with lysed cells and all three targeted proteins were identified in both whole cell and supernatant (Fig. 4A and 4B).
The nanobody-antigen interaction was verified by mixing intact E.coli cells displaying Neae-Nb3 with the supernatant of Ag3-eforRED (Fig. 3A). Red fluorescent characteristics were observed in the pellets after resuspending the centrifuged mixture, which is absent in the control group that only contains Neae-Nb3 (Fig. 4C).
After that, the type II cohesin-dockerin interaction was tested using the mixture of Neae-Nb3, OlpB-Ag3, and the type II dokerin fused with eforRED (Fig. 3B). A negative control lacking OlpB-Ag3 was set up for result comparison. Centrifugation was used to remove supernatant and the red fluorescence was only identified in pellets of the sample group, confirming the type II cohesin-dockerin interaction (Fig. 4D).
Finally, the association between type I cohesin and type I dockerin was validated using the mixture of Neae-Nb3, OlpB-Ag3, CipA1B2C, and DocI-eforRED (Fig. 3C), red fluorescence was detected in the resuspended mixture while it was not observed in the control group lacking the primary scaffold CipA1B2C (Fig. 4E), verifying the type I cohesin-dockerin interaction.
Cellulosome construction
We assembled the cellulose-like complex on the surface of E.coli by adding primary scaffold proteins, cellulases and cellulase boosters onto E.coli expressing secondary scaffold proteins. The mixture was centrifuged and resuspended in tris-HCl. The mixture underwent centrifugation and resuspension using tris-HCl, and cellulose was added to the mixture.
After 24h, the mixture was filtered and tested for glucose by Benedict's test. From the result, we determined that the cellulosome-like complexes are able to degrade cellulose at a higher efficiency than cell-free cellulases mixture. The overall success in engineering our project was verified by the successful construction of cellulosome complex and degrading cellulose to reducing sugars.

Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 314
Illegal XhoI site found at 1237 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 543
Illegal AgeI site found at 325
Illegal AgeI site found at 669
Illegal AgeI site found at 868
Illegal AgeI site found at 1539 - 1000COMPATIBLE WITH RFC[1000]
References
1. Anandharaj, Marimuthu et al. "Constructing A Yeast To Express The Largest Cellulosome Complex On The Cell Surface". Proceedings Of The National Academy Of Sciences, vol 117, no. 5, 2020, pp. 2385-2394. Proceedings Of The National Academy Of Sciences, https://doi.org/10.1073/pnas.1916529117.
| None |


